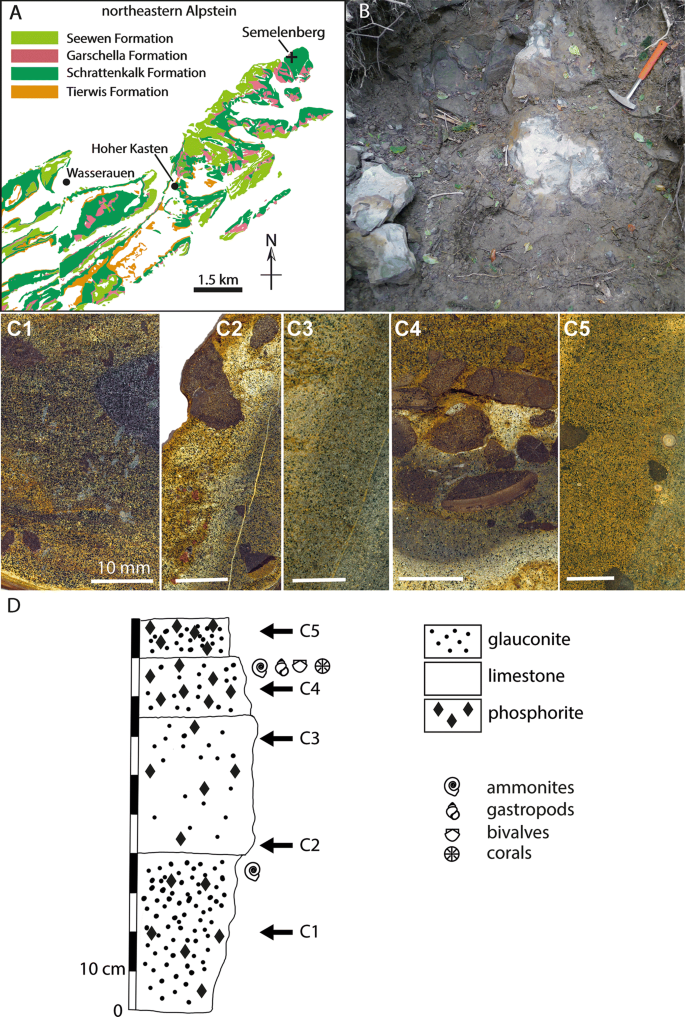
For 25 out of 112 targets, both scoring functions perform poorly, with BEDROC metrics lower than 0.1 (see Tables S2 and S3). Based on the BEDROC metric, the AD4 scoring function outperforms the Vina scoring function for the following targets: pur2, fpps, tryb1, xiap, and nram but underperforms for thb, fak1, ada17, hdac8, and pgh2.

However, the two scoring functions show different performance depending on the targets. The results ( Figure 3) show that overall both Vina and AD4 scoring function perform similarly in early recognition, but the Vina scoring function reproduces crystal poses with a higher accuracy. The correct pose is found in only one of the 10 docking replicates as the top 2 poses, and in only two replicates as the top 3 poses. However, a decrease in accuracy in observed for 4kyz, with an average RMSD of 5.2 Å for the top 1 pose using the hydrated docking, compared to an average RMSD of 1.9 Å without water molecules. For two of the six ligands (4yku and 4ykx), a net improvement is observed with RMSD below 1 Å compared to average RMSD values above 2 Å 6.1 and 2.3 Å for 4yku and 4ykx, respectively (see Table S1). Considering only the top pose, the success rate is increased by 17 percentage points, from 50 to 67%, and goes up to 83% with the top three poses. The results show that the docking success rate is higher using the hydrated docking protocol ( Figure 2).
The docking success rate is reported in Figure 2, and a hand-picked system (PDB 4ykq) is depicted in Figure 1B. Details about the screening library and receptor preparation and analysis method are discussed in Supporting Information. (28) This is an interesting system for the hydrated docking because different ligands bind with a different number of water molecules bridging hydrogen bonds with the protein. To validate the implementation of this docking protocol in Vina 1.2.0, we used six HSP90 protein–ligand complexes from the D3R Grand Challenge 2015. (14) Following the standard hydrated docking protocol, (14) the W map, which represents water–receptor interactions, is obtained by combining the oxygen-acceptor (OA) and hydrogen-donor (HD) maps of the AD4 force field. It is a constant value that was calibrated to 0.2 kcal/mol in the original study.
This entropy reward does not take the receptor into account. In fact, when that happens, a water molecule is considered displaced (i.e., removed from the system), and a reward is added to the score to reflect the entropy gain resulting from releasing the water molecule to the bulk solvent. During docking, W atoms move along with the ligand, do not contribute to intramolecular interactions, and are allowed to overlap with the protein. Water molecules are represented by a single atom of type W, and are added to the ligand molecule at the end of each hydrogen bond vector. (14) The method is based on docking ligands explicitly hydrated with spherical water molecules, and can be used to predict the position and the role (i.e., bridging or displaced) of individual water molecules and generally improve ligand pose predictions. The hydrated docking protocol was developed to model water molecules directly involved in the ligand–receptor interaction. It provides a more accurate representation of water molecule interactions with the receptor within ligand-binding sites, but at the expense of a much higher computational cost compared to implicit solvent models. (25) GIST is a method to analyze MD simulations and characterize thermodynamic properties of water molecules. The availability to accessible grid map files generated by AutoGrid provided the foundations for a number of specialized methods, such as the zinc-coordination potentials in the AutoDock4 Zn force field, (13) biasing docking using information from molecular dynamics (MD) simulations in AutoDock-Bias, (21) and the integration of Grid Inhomogenous Solvation Theory (GIST) (22−24) in AutoDock-GIST. In AD4, maps are precalculated using a separate program (AutoGrid (2)) prior to docking and loaded at runtime, while Vina calculates them on-the-fly prior to running the MC search. Vina also uses the target structure to perform a postprocessing minimization of the docked poses. Both the AD4 and Vina programs calculate intermolecular interactions by performing trilinear interpolations of grid maps precalculated on the target structure.
